Fear the superbug: Watch bacteria develop high-level drug-resistance in a giant Petri dish
The way the bacteria's evolution played out surprised the researchers in a couple of ways.
Crafty bacteria that mutate and evade the forces of current drugs are a real concern to the global community, with a recent report predicting that these superbugs could kill 10 million people a year by 2050, an average of one death every three seconds. Scientists have built a new tool to study how the microscopic killers operate, taking the form of giant Petri dish where they can be seen evolving resistance to rising concentrations of antibiotics in a relatively short space of time.
Much of the time, scientists study the evolution of microbes in the lab by pitting them against drug mixtures in relatively small spaces, like on tiny plates. Researchers at the Harvard School of Medicine (HMS) together with colleagues at the Technion-Israel Institute of Technology sought to create an environment that would more accurately mimic real-world conditions, allowing factors like space, size and geography to come into play.
"We know quite a bit about the internal defense mechanisms bacteria use to evade antibiotics but we don't really know much about their physical movements across space as they adapt to survive in different environments," says study first author and research fellow in systems biology at HMS, Michael Baym.
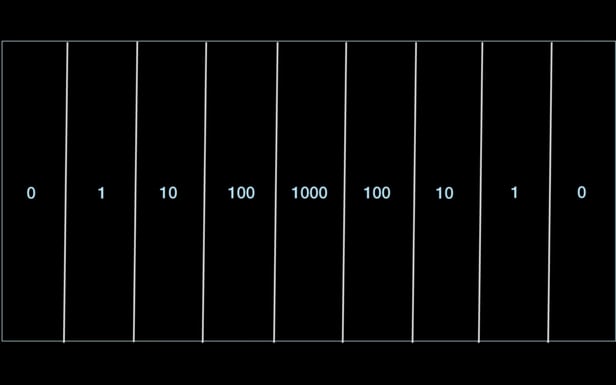
So the team built a huge Petri dish that they could use to observe bacterial behavior as the microbes encountered different concentrations of antibiotic drugs. Measuring 2 x 4 ft (0.6 x 1.2 m), the dish is split into nine different reservoirs running the length of the dish. All are filled with agar, a jelly-like substance taken from seaweed used in science experiments to nourish growing organisms, but each has differing concentrations of antibiotic drugs.
The sections at each of end contained no drugs at all, while the second from the end contained just above the minimum required to kill the bacteria. These dosages increased ten-fold in each section toward the center, with dosages of 10 times, 100 times and then 1,000 times more than needed to kill the bacteria.
By mounting a camera on the ceiling, the team created a time-lapse video (see below) that showcases evolution in all its glory. They introduced E. coli to the end sections and watched as the bacteria adapted, survived and thrived on each level until conquering the highly lethal dosage in the middle, standing triumphantly as superbugs capable of enduring a drug dosage 1,000 times stronger than that which killed their ancestors just 10 days earlier. When the scientists switched antibiotics, from trimethoprim to ciprofloxacin, the bacteria exhibited a 100,000-fold increase in drug resistance.
Dubbed the Microbial Evolution and Growth Arena (MEGA), the giant plate was designed as an educational tool so that students could see evolution playing out before their eyes, but the researchers found that it has revealed some useful insights about drug-resistant bacteria, too.
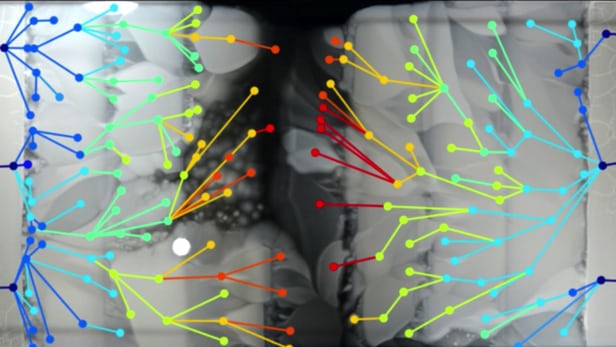
As the bacteria spread into new concentration levels, only a small portion were able to adapt and survive. Mutations through the generations would eventually give rise to drug-resistant microbes whose descendants then moved into new frontiers, and so-forth until they reached the center. But the way this played out surprised the researchers in a couple of ways.
They found that the initial mutations applied the brakes somewhat, with growth slowing as the bacteria adapted to each higher dose of antibiotics, before returning to normal growth rates once they had become fully resistant. Their observations also challenge the traditional assumption that the most resistant mutants are those responsible for driving evolution. By studying mutants both at and behind the front lines, the team found that this was not always the case.
"What we saw suggests that evolution is not always led by the most resistant mutants," says Baym. "Sometimes it favors the first to get there. The strongest mutants are, in fact, often moving behind more vulnerable strains. Who gets there first may be predicated on proximity rather than mutation strength."
While it doesn't perfectly replicate how bacteria behave in real-world environments, such as hospitals, the researchers say the MEGA-plate is a more accurate representation than typical lab cultures.
"Our MEGA-plate takes complex, often obscure, concepts in evolution, such as mutation selection, lineages, parallel evolution and clonal interference, and provides a visual seeing-is-believing demonstration of these otherwise vague ideas," says senior study investigator, Roy Kishony. "It's also a powerful illustration of how easy it is for bacteria to become resistant to antibiotics."
You can watch the bacteria take over the MEGA-plate in the video below, while the team's research was published in the journal Science.
Source: Harvard School of Medicine